humantumoratlas.org → NCI HTAN →
The Human Tumor Atlas Network (HTAN) is an NCI Cancer Moonshot initiative focused on building three-dimensional atlases of the cellular, morphological, and molecular features of human cancers as they evolve from precancerous lesions to advanced disease. KSG @ Dana-Farber serves as the Data Coordinating Center (DCC) alongside Sage Bionetworks, the Institute for Systems Biology, and Memorial Sloan Kettering Cancer Center.
What We Do
Manage and coordinate the submission, validation, and release of data across all HTAN research centers, ensuring completeness and consistency.
Develop and maintain the HTAN data model — a standardized metadata schema covering biospecimens, assays, clinical data, and files across all centers.
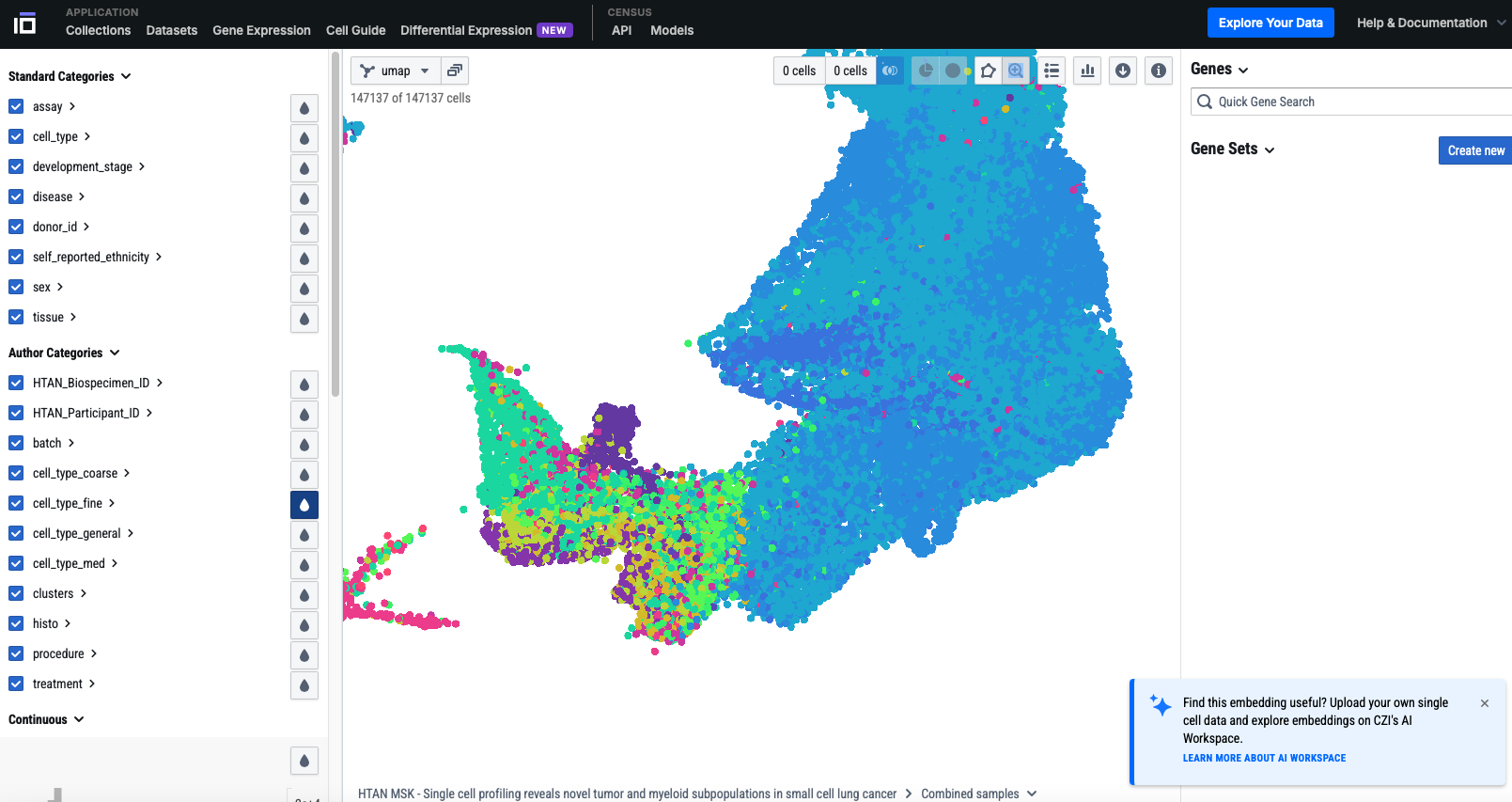
Build and maintain the HTAN Data Portal, enabling researchers worldwide to explore, filter, and download multi-omic datasets from participating centers.
Make HTAN data accessible via multiple cloud platforms — Synapse, Google BigQuery, and the NCI Cancer Research Data Commons (CRDC/Gen3) — for scalable analysis.
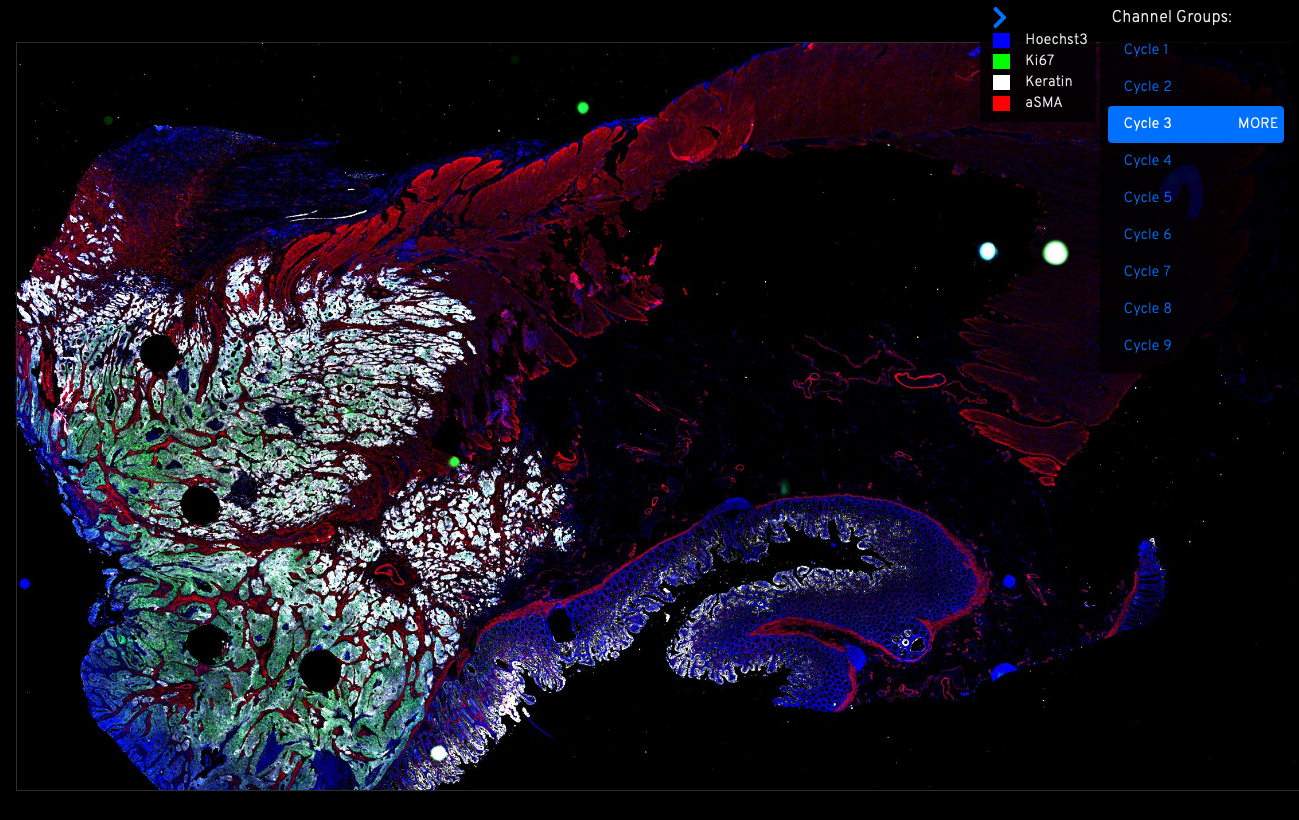
Support the collection and standardization of cutting-edge single-cell and spatially resolved datasets — including scRNA-seq, MIBI-TOF, CyCIF, and more.
Coordinate longitudinal cohorts tracking cancer evolution — from precancerous lesions through metastasis and drug resistance — with comprehensive clinical annotation.
Screenshots
About HTAN
HTAN brings together leading cancer research centers across the United States to study how cancers form, progress, and respond to treatment. Each HTAN center focuses on a specific cancer type or biological question — including colorectal cancer, breast cancer, lung cancer, melanoma, glioblastoma, and others — generating rich multi-omic datasets that are made openly available to the research community.
The network produces data from a wide range of technologies: bulk and single-cell RNA sequencing, whole-genome and targeted DNA sequencing, proteomics, imaging mass cytometry, multiplexed immunofluorescence, and spatial transcriptomics. All data is linked to clinical and pathological annotations and released under FAIR data principles.